RhoFold+: A Revolutionary Framework for RNA 3D Structure Prediction
RhoFold+: Unlocking the Secrets of RNA 3D Structures with Speed and Precision
RNA plays a crucial role in various biological processes, and understanding its 3D structure is key to unlocking insights into its function and potential applications in biotechnology. However, predicting RNA structures accurately has been a significant challenge—until now. Enter RhoFold+, a deep learning-based framework designed to predict RNA 3D structures with unprecedented speed and accuracy.
Key Features of RhoFold+
RNA Language Model
RhoFold+ employs a pre-trained RNA language model based on transformer architecture. This model, trained on 23 million sequences from 800,000 species, consists of 12 layers, making it highly robust for RNA structure prediction.Automated Pipeline
The framework is fully automated and does not require human intervention, simplifying the process of RNA structure prediction.Speed and Efficiency
RhoFold+ achieves an average inference time of just 0.14 seconds per sequence, making it one of the fastest tools in this domain.Secondary Structure Prediction
Beyond predicting 3D structures, it can also determine secondary structures and interhelical angles (IHAs), broadening its applications in RNA research.Generalization Capabilities
The model demonstrates strong generalizability by accurately predicting unseen RNA structures across diverse families and types.
Performance Highlights
RhoFold+ has outperformed state-of-the-art methods, including human expert-driven approaches, in benchmarks like RNA-Puzzles and CASP15. It achieves a mean root-mean-square deviation (r.m.s.d.) of less than 4 Å for non-overlapping datasets, showcasing exceptional accuracy. Additionally, it excels at predicting complex interactions and maintains robustness across various datasets.
Core Components of RhoFold+
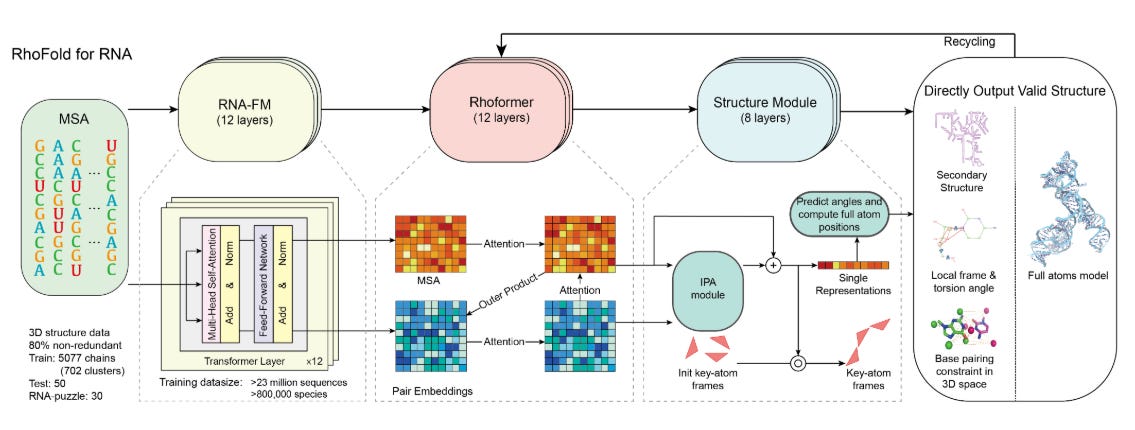
1. MSA (Multiple Sequence Alignment)
MSAs are generated using tools like Recursive MSA (rMSA) and Infernal, which search extensive sequence databases such as Rfam and RNAcentral.
These alignments capture conserved secondary structures essential for accurate predictions.
To optimize computational efficiency, RhoFold+ selects 256 MSAs either randomly or through clustering methods.
2. RNA Foundational Model
A BERT-like model pre-trained on 23 million RNA sequences serves as the foundation for feature extraction.
3. RhoFormer
The core transformer network of RhoFold+, RhoFormer refines embeddings over nine cycles.
It employs geometry-aware attention mechanisms to optimize local frame coordinates and Invariant Point Attention (IPA) modules to refine torsion angles.
4. Structure Module
This module uses geometric modeling techniques to predict 3D structures while enforcing biological constraints like base pairing.
Predictions are iteratively refined to improve spatial accuracy.
5. Post-Processing
Tools such as Assisted Model Building with Energy Refinement (AMBER) are used to resolve structural inaccuracies, ensuring biologically valid outputs.
Innovative Mechanisms: Geometry-Aware Attention & IPA
Geometry-Aware Attention
This mechanism enhances traditional transformer attention by incorporating spatial relationships among data points. It aligns tokens into a common coordinate space using geometric attributes like distances and angles, enabling better performance in tasks involving 3D molecular data.
Invariant Point Attention (IPA)
IPA refines torsion angles by focusing on intrinsic spatial relationships between nucleotides:
Operates on local frames parameterized by rotation matrices and translation vectors.
Iteratively updates nucleotide features based on geometric properties.
Ensures invariance to global transformations like rotation and translation.
Integrates locality-aware adjustments for handling RNA's structural flexibility.
Limitations
While RhoFold+ is groundbreaking, challenges remain:
Predicting large or highly flexible RNA structures (e.g., junctions and pseudoknots) is still difficult.
Further improvements are needed to address these complex cases effectively.
Why RhoFold+ Matters
RhoFold+ represents a significant leap forward in RNA structure prediction:
Its speed and efficiency make it suitable for high-throughput studies.
The ability to predict both secondary and tertiary structures broadens its utility in RNA engineering.
Its generalization capabilities open doors for studying previously unexplored RNA families.
With its innovative design and exceptional performance, RhoFold+ is set to transform the field of RNA research, paving the way for new discoveries in biology and medicine.
For Code implementation refer the github link : https://github.com/ml4bio/RhoFold