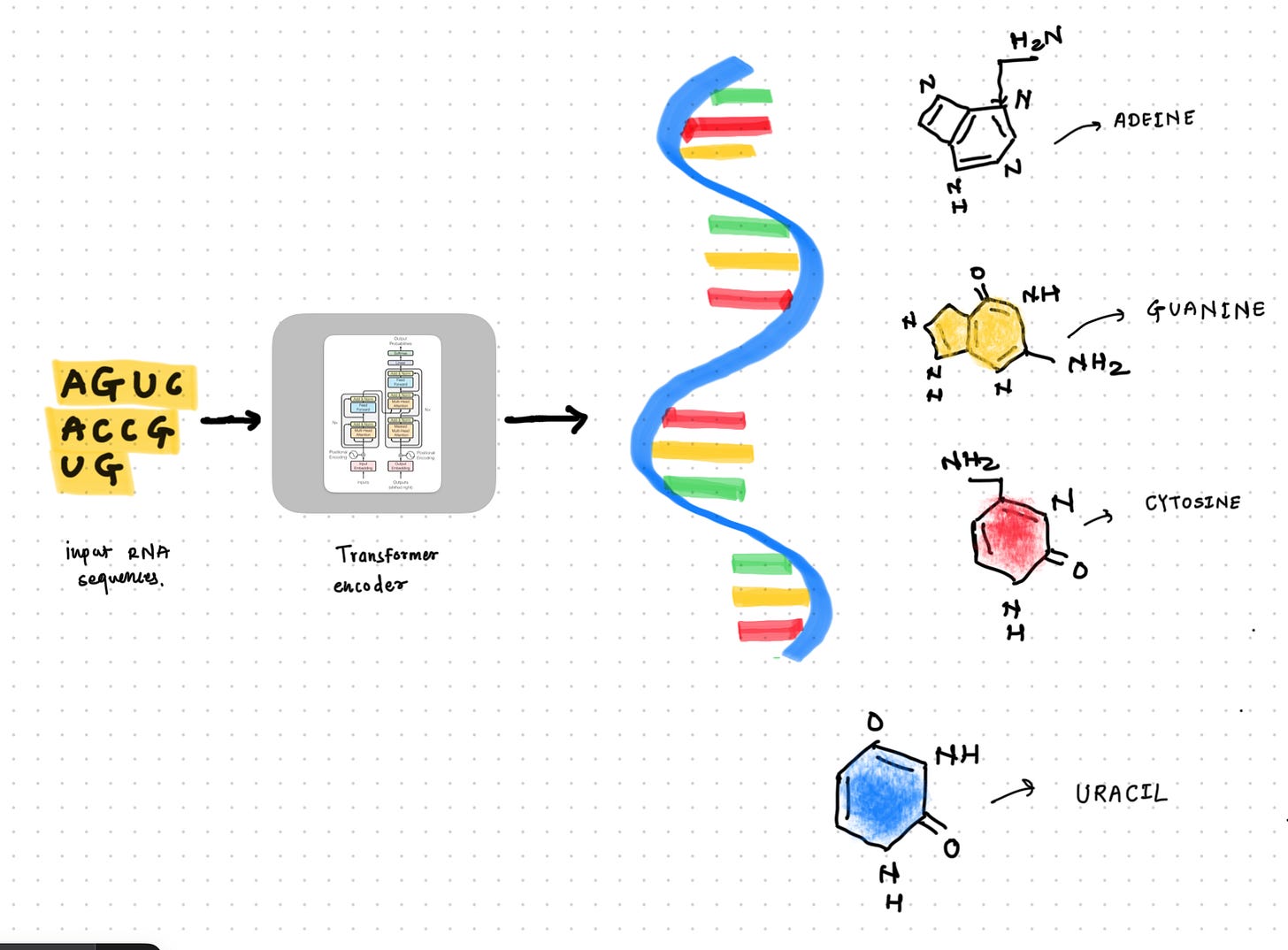
RNA Transformer
Model to encode the RNA sequences using transformer encoder architecture.
I joined the Kaggle competition Stanford RNA 3D Folding two months late. It was an out-of-domain competition for me, so I initially tried to understand the width of the domain by studying different models and techniques used to model the RNA sequences using Deep Learning. There are lots of interesting models to study, like ProteinX (a clone of AlphaFold3), RhoFold, and RibonanzaNet. I explored RibonanzaNet, Which is an Transformer encoder. I was interested to see what difference it makes to encode the RNA sequences compared to our generic transformer encoder which we use for English language modeling.
There are two major additions to the encoder architecture :
Outer product mean.
Triangular Multiplicative Module.
So, In this blog post we are exploring them.
1. Outer Product Mean :
Outer product mean used to learn the pairwise relationship between elements of the sequences. For example amino acids in proteins, residue in RNA etc.
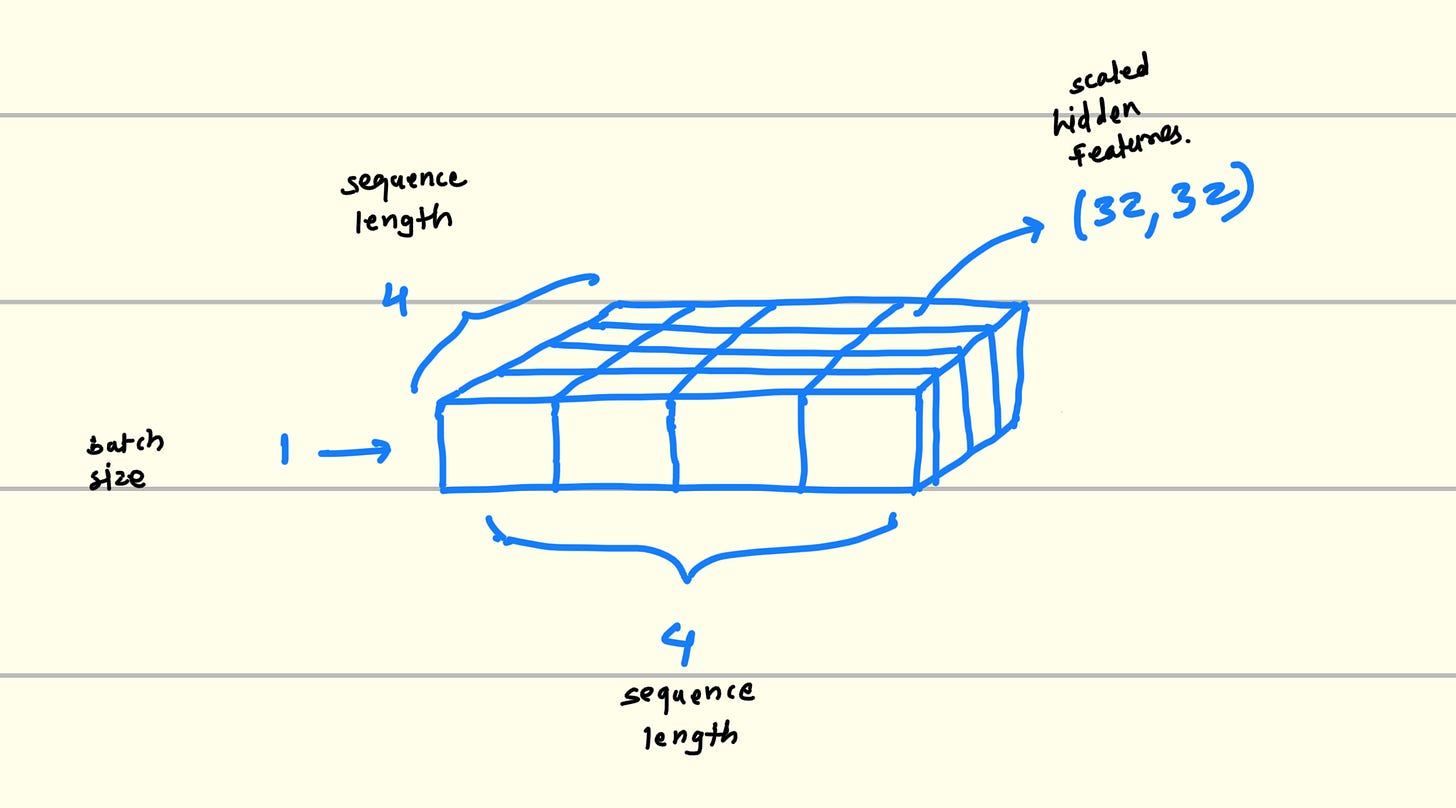
Technically, It creates one 2D matrix of seqence_length * seqence_length. where each (x, y) will give us one more 2D matrix of hidden_length * hidden_length which is actually highlights the patters if they have any.
This hidden_length * hidden_length matrix will have high value for those elements who have any relation.
What kind of possible relation or pattern can two residues have in between?
Complementarity : If some features from residue one is complementary to the another residue vector features then their corresponding values get highlighted in the 32*32 matrix.
Dependency : If the any two residue have the features which are dependent on each other then that 32*32 matrix in above diagram will have corresponding (x, y) elements highlighted.
1.1. Code :
class OuterProductMean(nn.Module):
def __init__(self, in_dim=256, dim_msa=32, pairwise_dim=64):
super(OuterProductMean, self).__init__()
self.proj_down1 = nn.Linear(in_dim, dim_msa)
self.proj_down2 = nn.Linear(dim_msa**2, pairwise_dim)
def forward(self, seq_rep):
seq_rep = self.proj_down1(seq_rep)
outer_product = torch.einsum('bid,bjc -> bijcd', seq_rep, seq_rep)
outer_product = rearrange(outer_product, 'b i j c d -> b i j (c d)')
outer_product = self.proj_down2(outer_product)
return outer_product1.2. Visualization :
1.3. Explanation :
Projecting features to lower dimension :
seq_rep = self.proj_down1(seq_rep)Projecting down the input embedding from 256 dimension to 32. It is just the linear transformation. from higher dimensions to the lower dimensions.
Outer product operation :
outer_product = torch.einsum('bid,bjc -> bijcd', seq_rep, seq_rep)Actually doing outer product in this step. here we create 32 * 32 matrix of feature vector which highlights patterns between pair of residues.
Flattening the pairwise feature vector :
outer_product = rearrange(outer_product, 'b i j c d -> b i j (c d)')Here, we are flattening vector on last dimension 32 * 32 into 1024.
Projecting features to lower dimension again :
outer_product = self.proj_down2(outer_product)
It just another linear layer to project features from higher dimensions to the lower dimensions.
2. Triangular Multiplicative Module
This is the key components in the state of art models like Alpha Fold.
This component is designed to enrich the pairwise representations by incorporating triangular relationships between RNA residues.
This allows to flow information through intermediate residue which is crucial for modeling non-local interactions and structural constraints in biomolecules.
There are two types for this
1. Outgoing flow of information.
2. Ingoing flow of information.
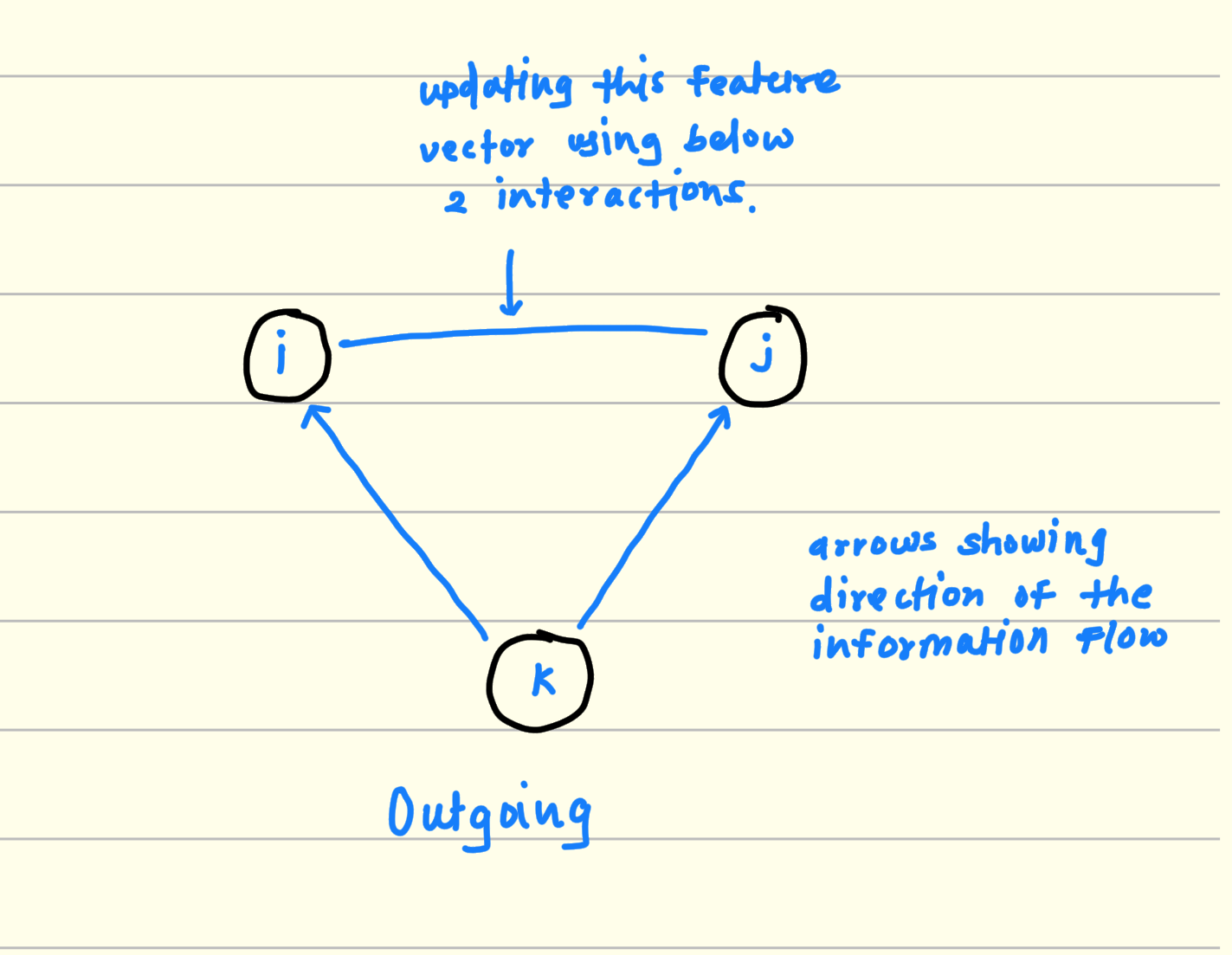
2.1 Outgoing flow of information
Pick any 3 residues. For example i, j, k.
Consider 2 sides from this triangle For example i, k and j, k.
multiply and aggregate the features from (i, k) and (j, k).
This output is the potential interaction between i and j considering k.
Keep iterating this process for all residue in the RNA sequence and finally you will get the triangular multiplicative interactions between RNA resides from the sequence for outgoing information flow.
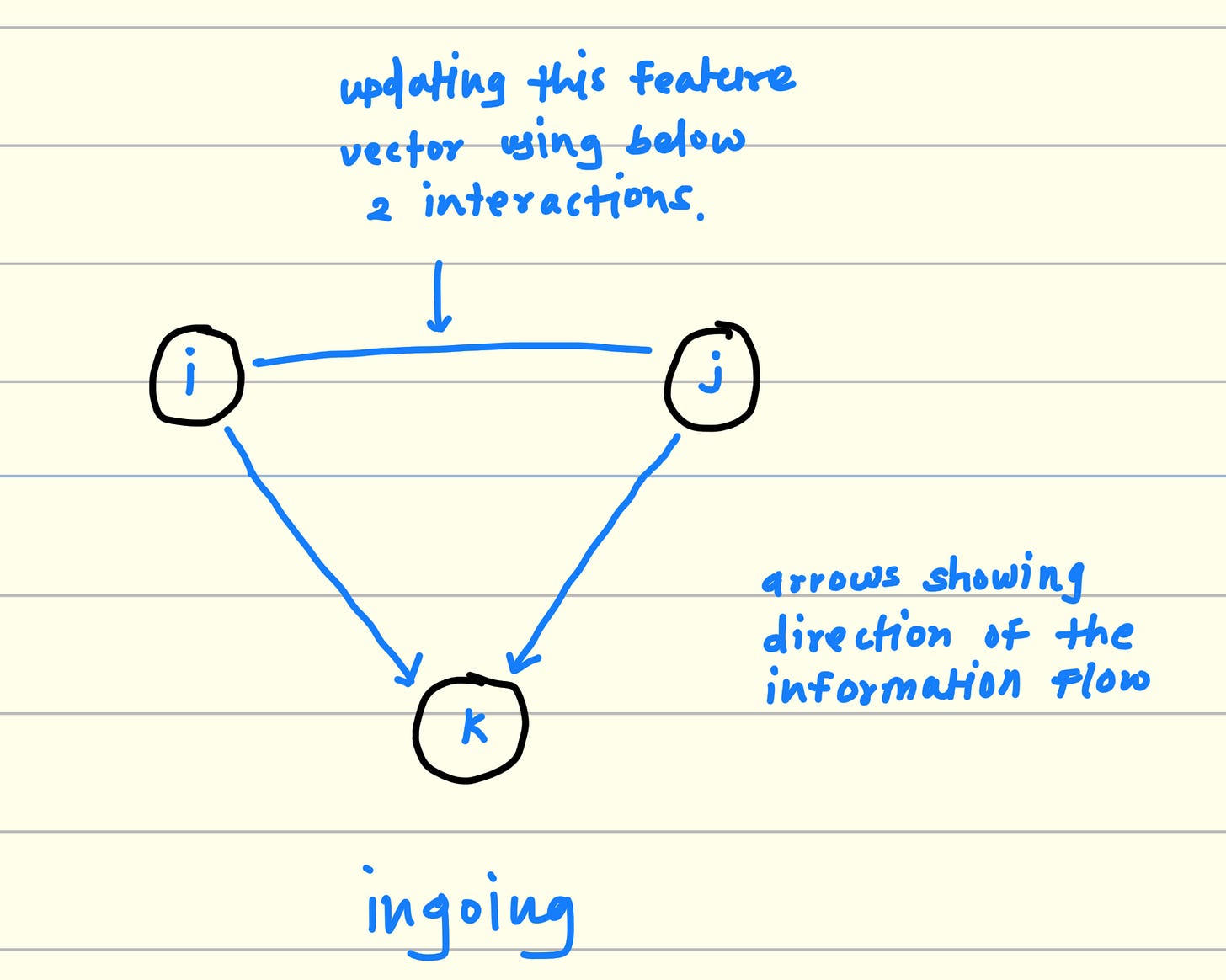
2.2. Ingoing flow of information
Again Pick any 3 residues form RNA sequence. For example i, j, k
Consider 2 sides from this triangle For example i, k and j, k
multiply and aggregate the features from (k, i) and (k, j).
This output is the potential interaction between i and j considering k.
Keep iterating this process for all residue in the RNA sequence and finally you will get the triangular multiplicative interactions between RNA resides from the sequence for ingoing information flow.
2.3. Code :
class TriangleMultiplicativeModule(nn.Module):
def __init__(self, *, dim, hidden_dim = None, mix = 'ingoing'):
super().__init__()
assert mix in {'ingoing', 'outgoing'}
hidden_dim = default(hidden_dim, dim)
self.norm = nn.LayerNorm(dim)
self.left_proj = nn.Linear(dim, hidden_dim)
self.right_proj = nn.Linear(dim, hidden_dim)
self.left_gate = nn.Linear(dim, hidden_dim)
self.right_gate = nn.Linear(dim, hidden_dim)
self.out_gate = nn.Linear(dim, hidden_dim)
# initialize all gating to be identity
for gate in (self.left_gate, self.right_gate, self.out_gate):
nn.init.constant_(gate.weight, 0.)
nn.init.constant_(gate.bias, 1.)
if mix == 'outgoing':
self.mix_einsum_eq = '... i k d, ... j k d -> ... i j d'
elif mix == 'ingoing':
self.mix_einsum_eq = '... k j d, ... k i d -> ... i j d'
self.to_out_norm = nn.LayerNorm(hidden_dim)
self.to_out = nn.Linear(hidden_dim, dim)
def forward(self, x, src_mask):
if exists(src_mask):
src_mask = src_mask.unsqueeze(-1).float()
mask = torch.matmul(src_mask,src_mask.permute(0,2,1))
assert x.shape[1] == x.shape[2], 'feature map must be symmetrical'
if exists(src_mask):
mask = rearrange(mask, 'b i j -> b i j ()')
x = self.norm(x)
left = self.left_proj(x)
right = self.right_proj(x)
if exists(src_mask):
left = left * mask
right = right * mask
left_gate = self.left_gate(x).sigmoid()
right_gate = self.right_gate(x).sigmoid()
out_gate = self.out_gate(x).sigmoid()
left = left * left_gate
right = right * right_gate
out = einsum(self.mix_einsum_eq, left, right)
out = self.to_out_norm(out)
out = out * out_gate
return self.to_out(out)
2.3.1 Linear layer’s
In the above code all linear layers do the work of lower projecting the dimensions form dim → hidden_dim, like if we have inputs of shape (1,6,6,64) and our dim is 64 and hidden dim is 32 then their output will be the matrix of shape (1,6,6,32).
As we are using these linear layers in multiple following mathematical operations in the process of model training their purpose becomes unique while model training.
Like proj layer just project the dimension from higher to lower and gates are adding non linearity to the information flow.
2.3.2 einsum
This is the core of the triangular multiplicative module. It is the Einstein summation function we can use for any matrix operation. It looks cleaner and it is optimized.
To code the outgoing and ingoing information flow we are using this functions. I have tried visualizing the function internal working in the following diagrams.
out = einsum(self.mix_einsum_eq, left, right)Before going into the depth of the einsum summation equation first we have to understand the matrix multiplication of 2 matrix each of shape (1,64) from the following diagram.
above equation shows the matrix multiplication of two matrices having same shape. it is too easy and basic thing from mathematics.
2.4. Visualization of Outgoing flow of information calculation :
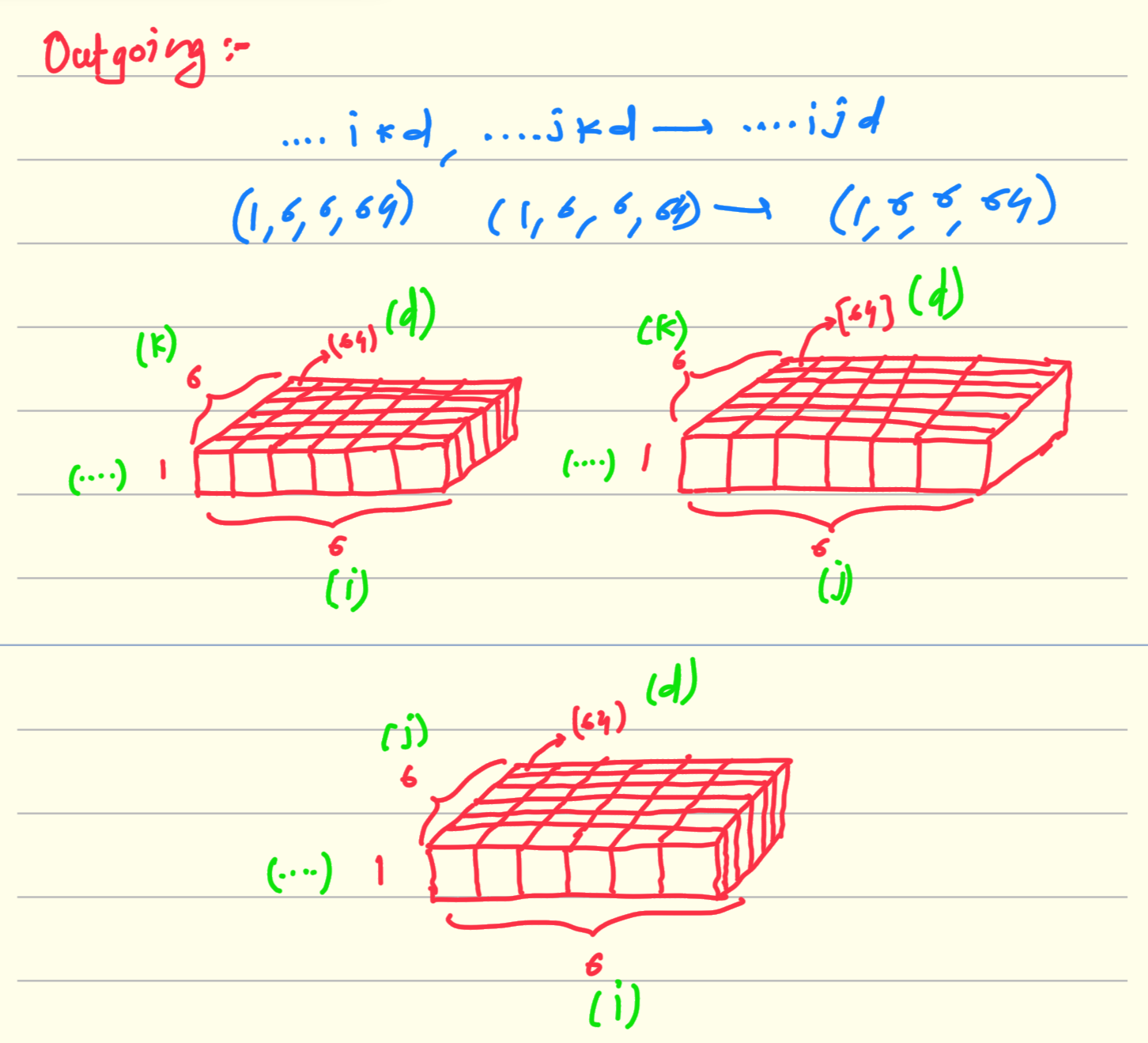
Einstein equation : ….ikd, …jkd → …ijd
It is the rule that the dimension which are not mentioned in the output of the above equation will get multiplied and aggregated. in above equation it is k. So multiplication of above two matrices of shape (1,6,6,64) results in matrix of same shape.
There are 36 matrices of shape (1,64) in left as well as in right matrix. we takes one from each and just do the matrix multiplication shown in above diagram.
Ingoing :
Einstein equation : ….kjd, …kid → …ijd
It is same operation as above the difference is that we are just changing dimension by which we are doing aggregate and multiply. In ingoing operation we are doing it on 2nd dimension from shape (1,6,6,64) and previously for outgoing information flow we did it on 3rd dimension form the same matrix. which is 6 in both the cases.
By just changing the dimension by which we have to do multiply and aggregate gives us the working of triangular multiplicative module. It is the beauty of matrix operations and specifically “Einstein summation notation“ which keeps the code cleaner and beautiful.
Conclusion :
We have studied Outer product mean and Triangular multiplicative module from the RNA model. Conceptually and it’s code implementation. If you still have any doubt reach out to me.
Whole encoder model code is here. GitHub repo : RNATransFormer. Except above two modules other things are same from the transformer encoder module described in the Attention Is All You Need paper.
References :
https://github.com/Shujun-He/RibonanzaNet.
https://www.kaggle.com/competitions/stanford-rna-3d-folding.